A junior radiologist is on call, scrolling through breast MRI slices at midnight.
On the second monitor, a segmentation mask, the tumor neatly outlined in electric blue, flickers into place, courtesy of an AI model trained and “validated” on one of the field’s best-known benchmarks.
She trusts it more than she admits. The benchmark scores were excellent. Papers said so.
None of those papers mention that for a woman ten years younger than the average in that dataset, the AI is not just a little worse. It’s systematically wrong in ways the benchmark can’t even see.
TL;DR
- Automated labeling bias turns benchmarks into a biased ruler: models agree with bad labels and look “fair” while failing real patients.
- In breast MRI, younger patients don’t just have noisier labels, they have qualitatively harder tumors, so training on automated labels can amplify age bias by ~40%.
- This isn’t ordinary label noise; it’s a measurement failure that changes how we should read medical AI benchmark scores, right now.
Automated labeling bias: the “biased ruler” that hides harm
In the MAMA‑MIA breast DCE‑MRI dataset, many tumor masks weren’t drawn by humans; they came from an automated pipeline. That’s the kind of shortcut radiology is rushing toward everywhere, foundation models sketch the outlines, humans maybe spot‑check a few.
Parikh et al. go back and audit those labels by age group. They find what you’d expect from years of fairness work: segmentation models do worse on younger patients. But when they evaluate models using those same automated labels as ground truth, the measured age gap shrinks or disappears.
That’s the biased ruler: if your measuring stick is warped, everything you measure looks straight.
The trick is subtle. Because models are trained and evaluated on the same family of machine‑generated labels, their predictions line up with the labels’ quirks. Benchmark metrics, Dice scores, IoUs, shoot up, including on the subgroups that are actually getting the worst real‑world performance.
From the outside, nothing looks wrong. From the inside, the model and the labels are quietly agreeing on the same mistakes.
And the field is sleepwalking into this pattern. We’ve already written about automated labeling bias in “AI builds AI” pipelines and the AI content feedback loop. Medical imaging is adopting the same playbook, just with tumors instead of text.
Younger patients break segmentation models, it’s qualitative, not just noisy
The comforting story would be: “Younger breasts are denser, so the labels are a bit messier. Clean up the noise and fairness improves.”
The MAMA‑MIA audit tears that up.
Parikh et al. take the standard excuses, label noise, “a few hard cases,” imbalanced difficulty, and test them systematically. They:
- Compare automated labels to expert references to see where errors cluster.
- Reweight and rebalance the training data by case difficulty.
- Probe whether models simply overfit to easier, older cases.
If the problem were just noisier labels for younger patients, you’d expect:
- Models to be unusually sensitive to those label errors.
- Fairness gaps to close when you reweight by difficulty or fix the ugliest labels.
Instead, the gap persists. Balancing by difficulty doesn’t help. Automated labels for younger patients are worse, yes, but even when you control for that, segmentation models still fall apart more often on younger cases.
The tumors themselves differ.
They’re larger, more variable in shape, and embedded in backgrounds that look less like the “canonical” cases the model has seen. It’s not “more of the same” difficulty; it’s different kinds of anatomy and pathology, a qualitatively different distribution.
That distinction matters. If age bias were mostly about noisy labels, you could treat it as a cleaning problem. But if the morphology is different, fairness is a representation problem: the model simply doesn’t have the right vocabulary of shapes and appearances to do the job.
Automated labels are built from that same defective vocabulary.
Automation can amplify bias: what the MAMA‑MIA audit shows
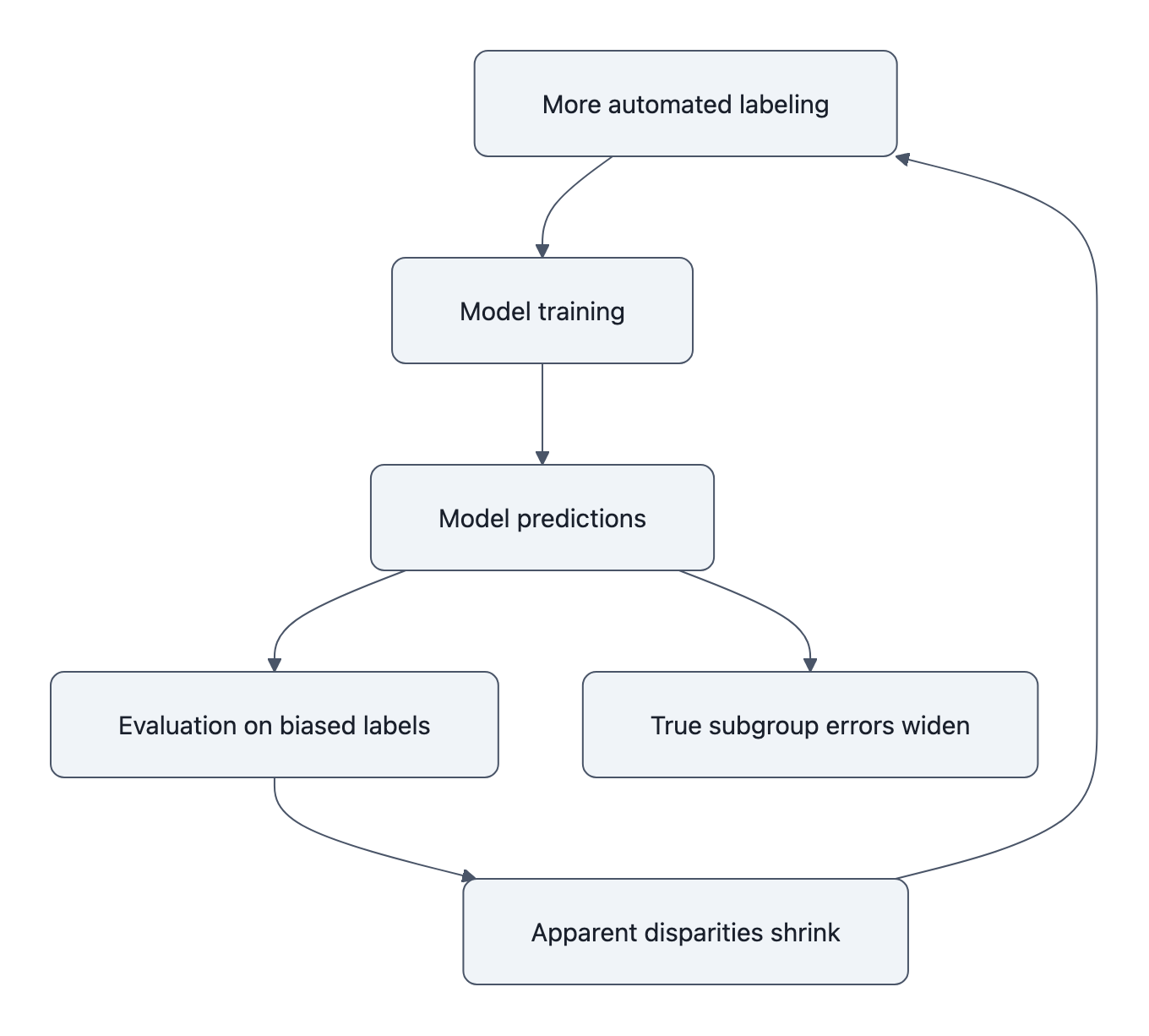

Here’s where the story gets stranger.
You might think: “Okay, the labels are biased. But at worst, models will inherit that bias, not go beyond it.”
The MAMA‑MIA audit finds something nastier: when you train segmentation models directly on the biased automated labels, age disparities grow, by on the order of 40% in their experiments, compared to training on cleaner human annotations.
The pipeline looks innocent:
- Use a strong model (or ensemble) to auto‑segment tumors in a large dataset.
- Treat those as “ground truth” labels, maybe with cursory checks.
- Train a new model on this big, shiny dataset.
- Evaluate it on a test set that is labeled… in exactly the same way.
What can go wrong?
Under the hood, automated labels tend to be most accurate on the majority, “easy” subgroup, the older patients with textbook tumor morphology. They’re least reliable on the atypical, harder distributions, the younger patients with weird, jagged lesions.
So the new model gets:
- Extremely consistent training signal on older cases (lots of similar shapes, all reinforced).
- Inconsistent, sometimes outright wrong signals on younger cases (shapes that don’t fit the learned template, labels that occasionally lie).
Standard optimization does the obvious thing: it fits the clean, abundant pattern and treats everything else as noise. That’s how bias amplifies.
Then the biased ruler comes back. You evaluate the biased model on a biased test set:
- It agrees beautifully with the automated labels on older patients, high scores!
- It agrees more often with the labels’ errors on younger patients, still decent scores!
The benchmark reports narrow gaps or none at all. Underneath, the model’s actual performance divergence has widened.
If you’ve read the classic “pervasive label errors destabilize benchmarks” paper, this is that idea with a twist. The problem isn’t just that wrong labels shuffle model rankings. It’s that systematic, subgroup‑skewed wrongness lets a whole class of models look good while quietly hurting the same people in the same way.
Why this automated labeling bias matters now
None of this is hypothetical.
MAMA‑MIA is a real, multi‑center dataset, released with the best intentions as a public resource. It joins a growing list of benchmarks that everyone cites and optimizes for, and that audits later reveal to be lopsided in who they serve well.
The Nature Medicine review on fair medical imaging AI is blunt: models that ace internal benchmarks frequently stumble on new hospitals, new scanners, new demographics. We add MAMA‑MIA’s twist: even before you leave the dataset, your benchmark can lie to you about subgroup harms.
That changes how we should read every “state‑of‑the‑art” segmentation score built on automated labels:
- A single metric, averaged over all patients, is no longer reassuring; it might just reflect how well you’ve overfit the majority’s label quirks.
- Even subgroup metrics can be suspect if the subgroups share the same automated labeling pipeline. They’re all measured with the same biased ruler.
- Small fairness gaps in benchmark tables aren’t evidence of safety; they might be evidence of shared delusion between model and labels.
It also changes what “scaling up” means. Automated annotation has been sold as a neutral tradeoff: a little noise in exchange for a lot more data. The MAMA‑MIA audits say: not neutral. When the noise is structured by age, ethnicity, site, by who is already underserved, more data can mean more damage, and better‑looking numbers.
This is the kind of failure mode that doesn’t announce itself. The models train. The curves go up. The reviewers nod at the Dice scores and accept the paper.
Meanwhile, the junior radiologist at midnight is looking at a clean blue contour around the wrong part of a young woman’s tumor, and the system that produced it has the graphs to “prove” it’s doing fine.
Key Takeaways
- Automated labeling bias turns benchmarks into a biased ruler: training and testing on the same flawed labels makes unfair models look fair.
- In breast MRI, younger patients have qualitatively different tumors, not just noisier labels, so standard fixes like difficulty balancing don’t close the gap.
- Training on machine‑generated labels can amplify age disparities beyond what was present in human annotations, even as benchmark metrics improve.
- Benchmark scores built on automated labels should be treated as suspect measurements, not ground truth, especially for subgroup performance.
- Fair medical imaging AI will require investing in clean, subgroup‑aware evaluation labels, not just bigger datasets with cheaper automated masks.
Further Reading
- Investigating Label Bias and Representational Sources of Age-Related Disparities in Medical Segmentation, Parikh et al.’s primary audit of MAMA‑MIA, introducing the “biased ruler” and showing bias amplification from automated labels.
- Who Does Your Algorithm Fail? Investigating Age and Ethnic Bias in the MAMA‑MIA Dataset, Companion preprint dissecting age, ethnicity, and site‑specific failures hidden by automated segmentation labels.
- MAMA‑MIA dataset paper, Original description of the breast DCE‑MRI dataset and benchmark that later audits scrutinize.
- Pervasive Label Errors in Test Sets Destabilize Machine Learning Benchmarks, Foundational evidence that label errors can flip benchmark conclusions, setting the stage for why biased “ground truth” is dangerous.
- The limits of fair medical imaging AI in real‑world generalization, Nature Medicine review on why high benchmark performance often fails to translate into fair, reliable clinical outcomes.
The next time you see a medical AI touting its benchmark score, imagine that blue contour on the midnight screen and ask a quieter question: what ruler did they use to measure this, and who did it bend against?